Schedule
-

3:00 PM
Introduction, goals, scope
-

3:10 PM
DICOM: Overview and historical perspective - David Clunie
This talk will introduce the main concepts of the DICOM standard, the role it plays in the modern healthcare enterprise, its overall purpose and capabilities.
-

3:40 PM
DICOM for Quantitative Imaging - Andrey Fedorov
This talk will discuss the relevance of the DICOM standard in quantitative imaging research, motivate its use and discuss specific capabilities as applied to the storage of common image analysis outputs, such as image segmentation results, parametric maps and region-based quantitative measurements.
-

16:05 PM
DICOM Data Management Systems - Marco Nolden
An overview, capabilities and summary of experience with the various data management platforms that can work with the DICOM data for medical image computing applications. Some of the tools we plan to cover will include XNAT and dcm4chee.
-

4:30 PM
Break
-

5:00 PM
Hands-on tutorial prerequisites setup
Attendees will be guided over the steps of installing the free open source software tools that will be used in the tutorial, and which can help the attendees work with the DICOM data in their everyday work after the conference. The tools utilized in the tutorial will include 3D Slicer, MITK, DCMTK, Atom, and dcmqi. We will also distribute the sample dataset that will be used in the tutorial.
Closer to the event we will provide instructions on how to install the prerequisites for your platform in advance of the tutorial! -
5:30 PM
Hands-on session: Using DICOM to store your analysis results
In this hands-on exercise attendees will use one or more open source tools (3D Slicer, MITK, dcmqi) to load DICOM data, segment regions of interest, and extract radiomics features. The results will be stored using standard (no private attributes) DICOM objects. The resulting standard-compliant data storage will be contrasted with the alternative approaches used in the community.
-
6:15 PM
Hands-on session: DICOM data wrangling
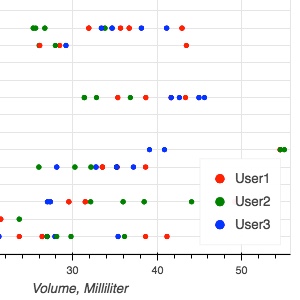
In this session we will go over the steps of extracting various data attributes (both in the source images, and in the research results stored in DICOM representation) from the standard DICOM representations of the analysis results, generating summary visualizations and preparing representations (e.g., CSV spreadsheets) suitable for the analysis by non-DICOM-enabled tools.